Research Articles
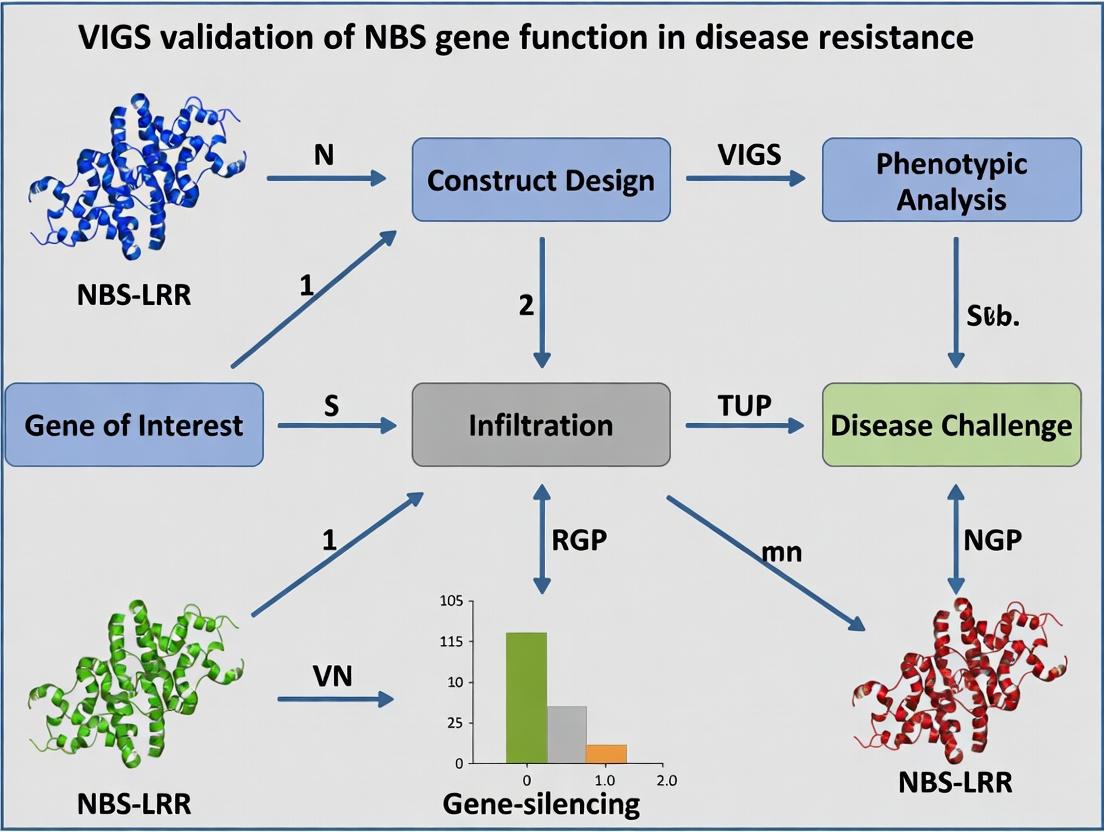
VIGS Validation of NBS-LRR Gene Function: A Comprehensive Guide for Disease Resistance Research
This article provides a detailed framework for using Virus-Induced Gene Silencing (VIGS) to validate the function of Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) genes in plant disease resistance pathways.
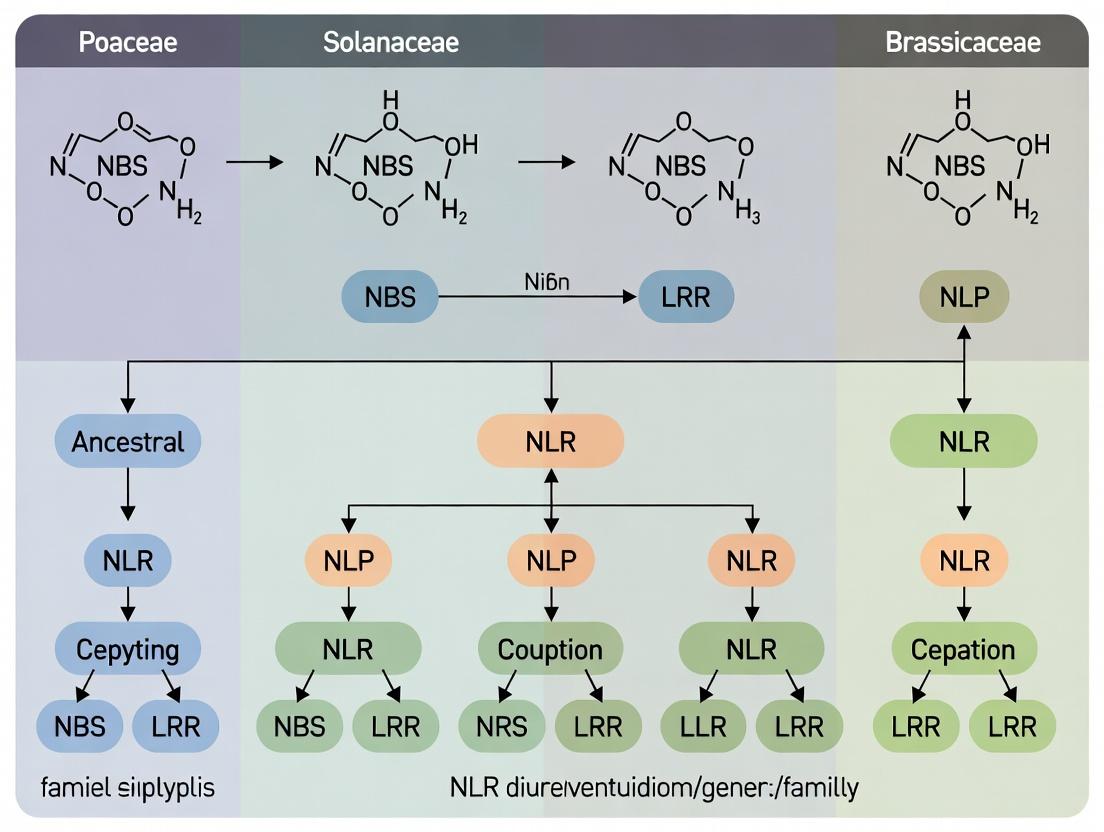
Evolutionary Dynamics of NLR Genes: Conservation Mechanisms and Family-Specific Diversification in Plant Immunity
This article provides a comprehensive analysis of NLR (Nucleotide-Binding Leucine-Rich Repeat) gene evolution across plant families.
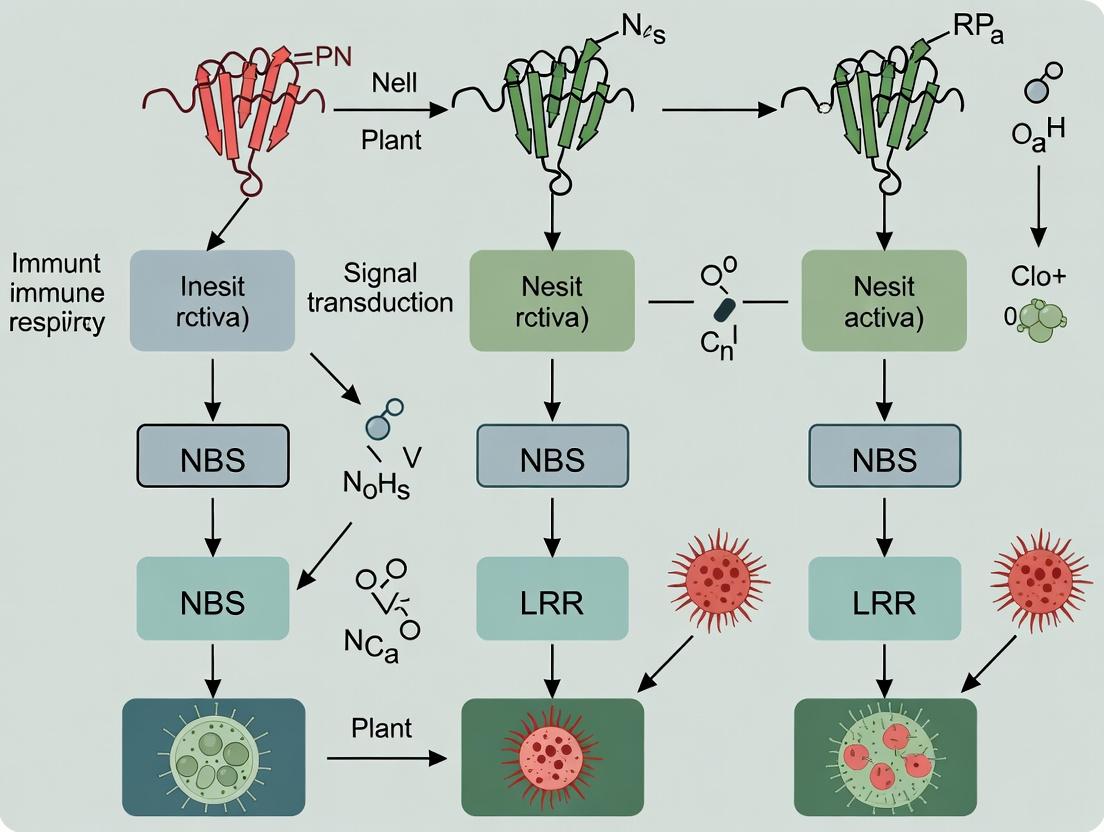
Guardians of Green Medicine: How NBS-LRR Genes Drive Disease Resistance in Medicinal Plants
This article provides a comprehensive exploration of Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) genes as central players in disease resistance within medicinal plants.
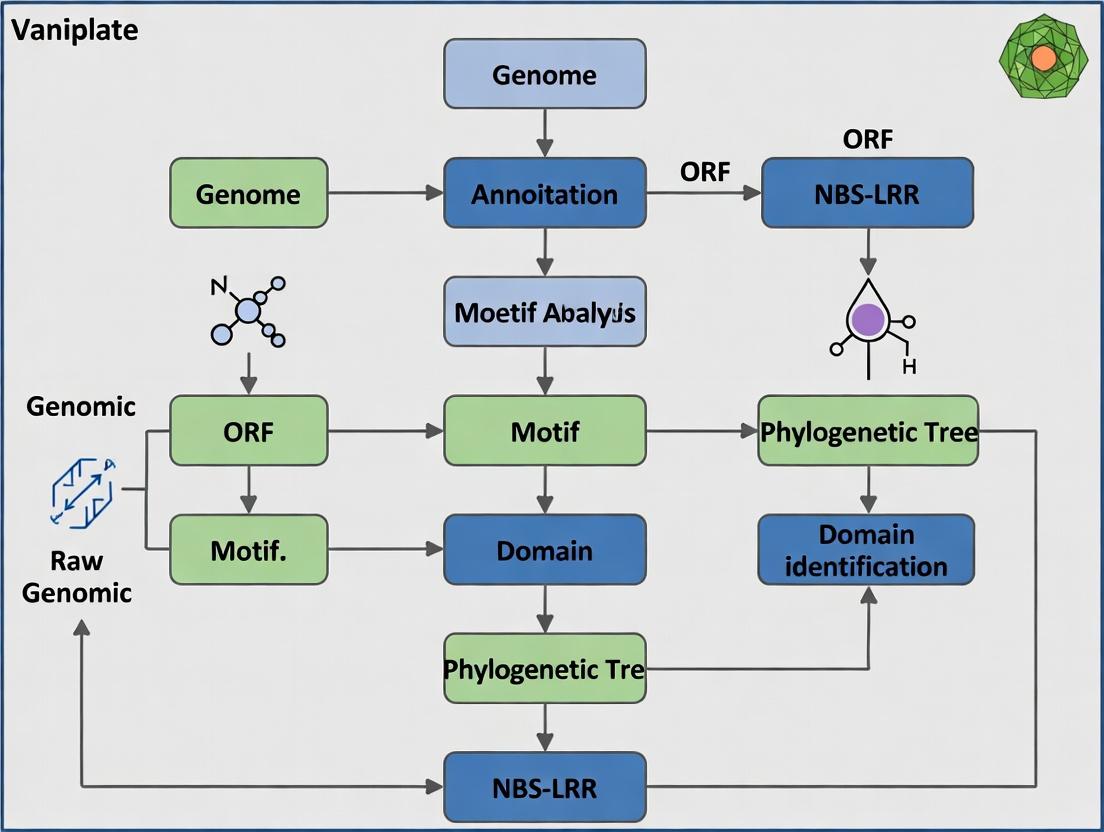
Genome-Wide Identification and Functional Analysis of NBS-LRR Genes in Salvia miltiorrhiza: Implications for Disease Resistance and Bioactive Compound Production
This comprehensive study provides a detailed genome-wide analysis of the NBS-LRR (Nucleotide-Binding Site-Leucine-Rich Repeat) gene family in the medicinal plant Salvia miltiorrhiza (Danshen).
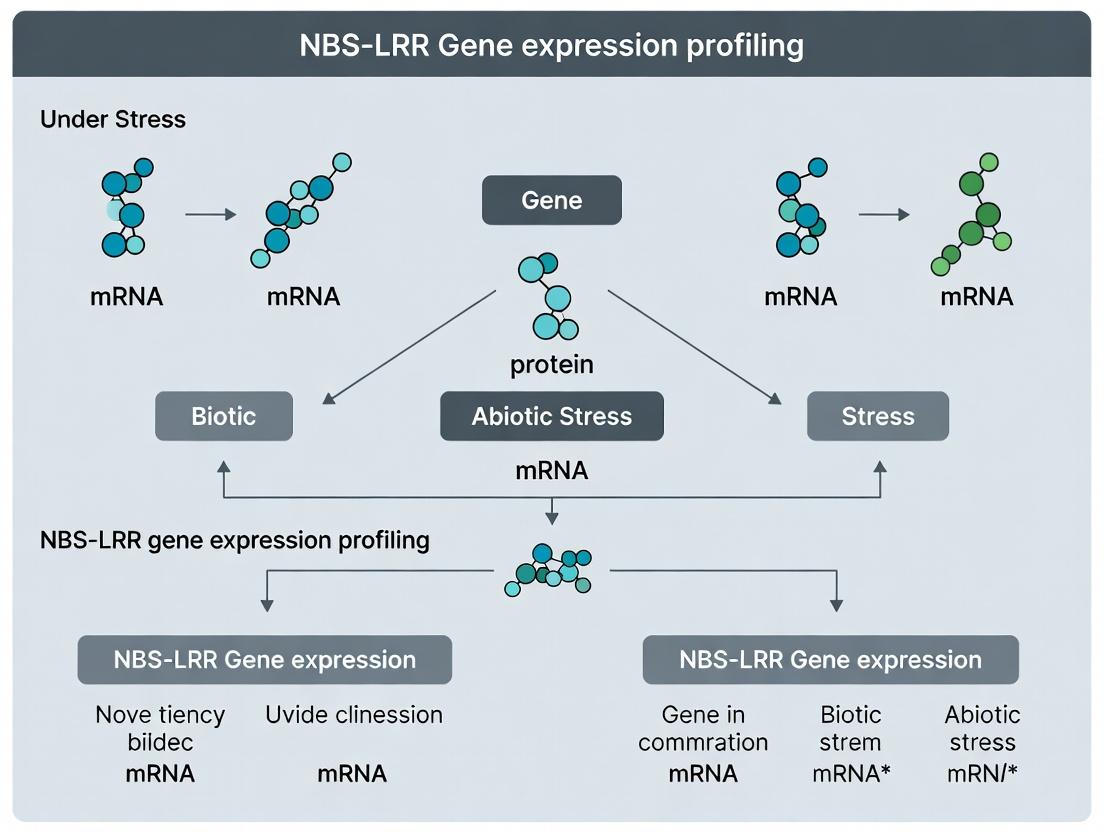
Decoding Plant Defense: A Comprehensive Guide to NBS-LRR Gene Expression Analysis Under Biotic and Abiotic Stress
This article provides a detailed methodological and analytical framework for profiling NBS-LRR gene expression in plants under stress conditions.
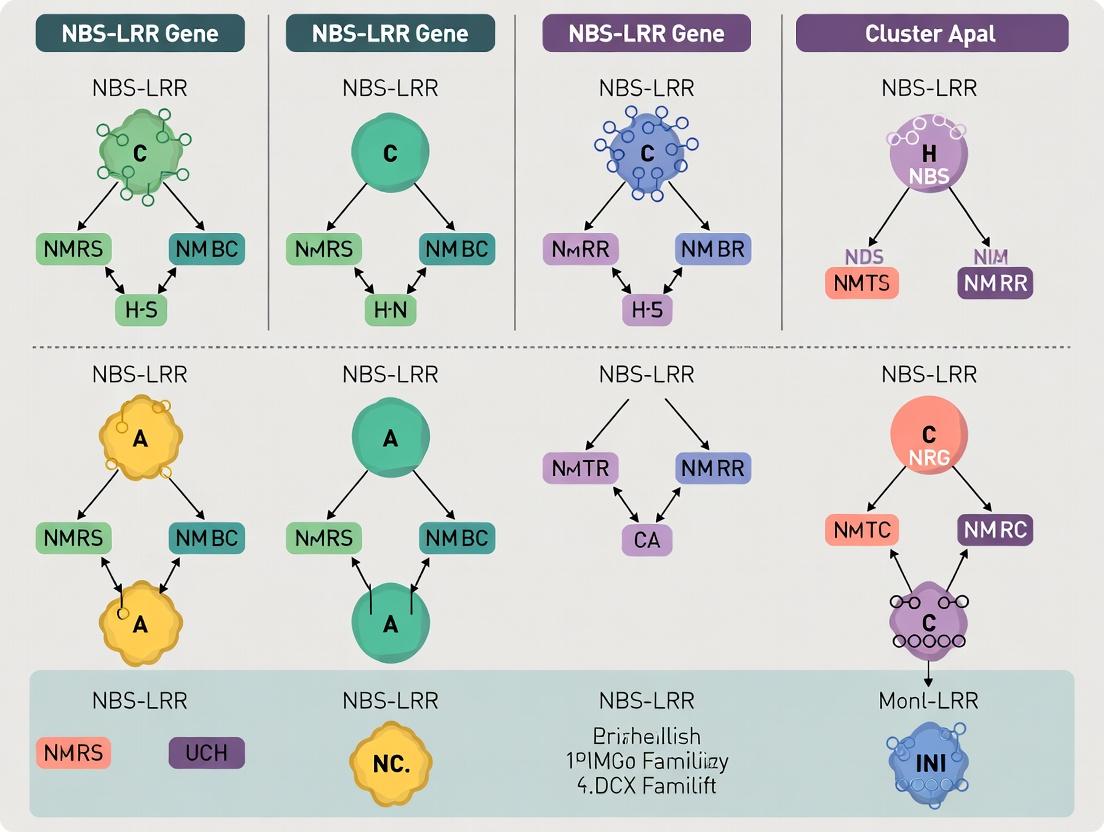
Decoding Plant Immunity: A Comprehensive Guide to NBS-LRR Gene Distribution and Cluster Analysis for Drug Discovery
This article provides researchers, scientists, and drug development professionals with a structured analysis of NBS-LRR genes, the cornerstone of plant innate immunity.
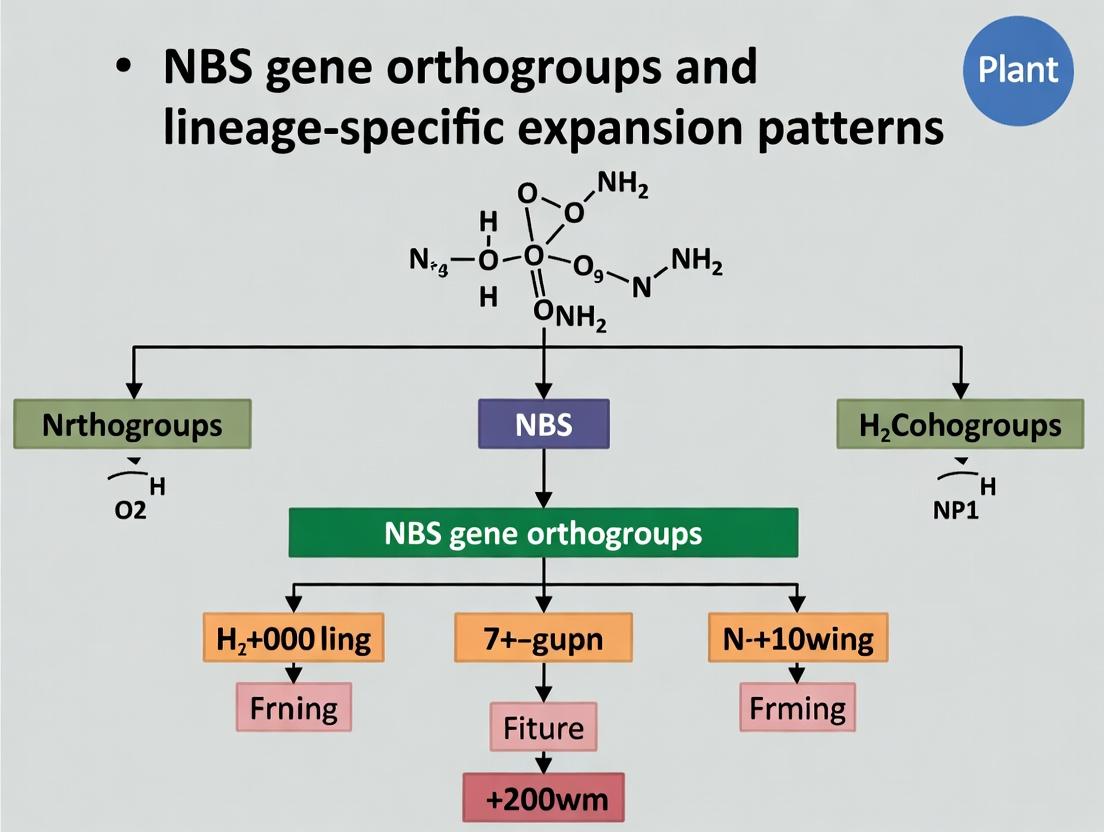
Evolutionary Dynamics of NBS Gene Orthogroups: Decoding Lineage-Specific Expansion for Biomedical Discovery
This article provides a comprehensive analysis of Nucleotide-Binding Site (NBS) gene orthogroups and their lineage-specific expansions, a critical area in plant-pathogen interaction genomics and disease resistance research.
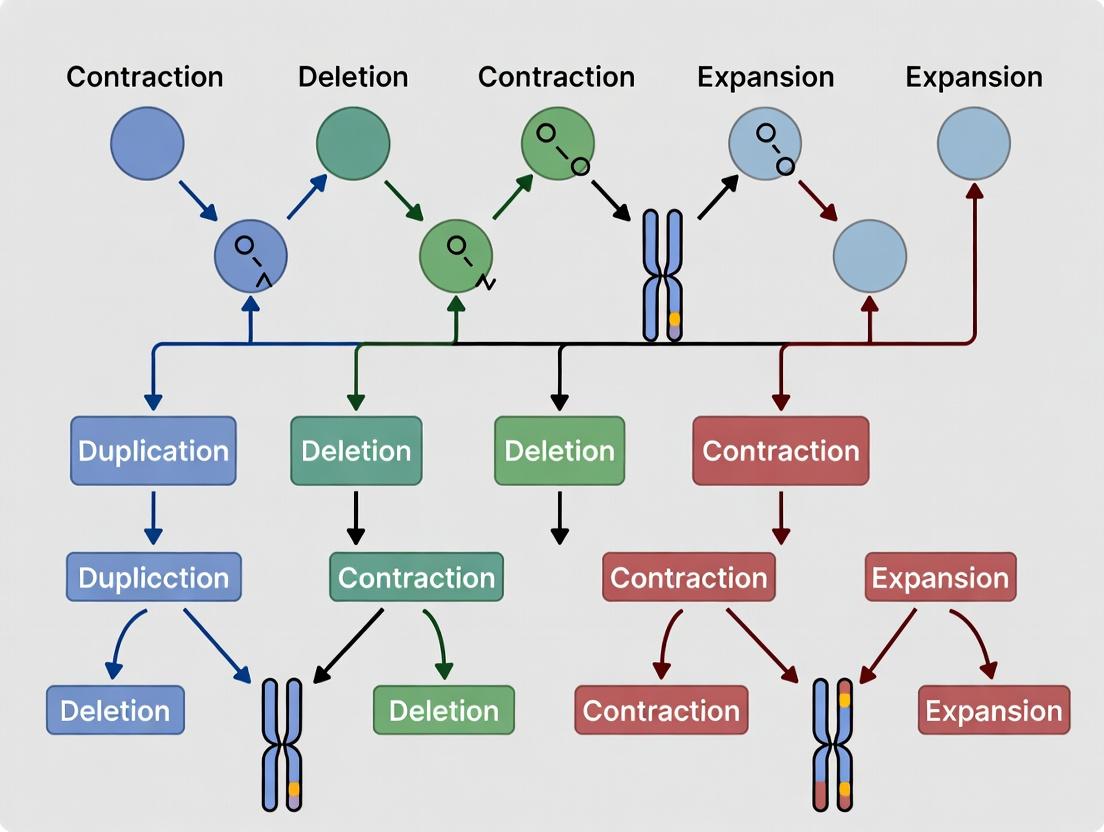
The NBS Gene Family: Unraveling Evolutionary Patterns of Contraction and Expansion in Plant Immunity
This article explores the dynamic evolutionary patterns of Nucleotide-Binding Site (NBS) gene families, key players in plant innate immunity.
Duplicating Defense: How Whole-Genome and Tandem Duplications Expand the NBS Gene Family in Plant Immunity
This article provides a comprehensive analysis of NBS (Nucleotide-Binding Site) gene expansion mechanisms, focusing on whole-genome duplication (WGD) and tandem duplication.
NBS Gene Diversification in Land Plants: Evolution, Disease Resistance Pathways, and Biomedical Research Implications
This comprehensive review explores the evolutionary diversification of Nucleotide-Binding Site (NBS) genes across land plants and its critical implications for disease resistance.